- Blog
- How to use family tree maker 2014
- Copytrans manager for mac download
- Hello neighbor act 1
- Eenie meenie miney mo twd
- How to play online dragon ball unreal
- Age of empires 3 scenarios
- Lawrence welk resorts escondido california
- 2caudio aether download
- Fl studio autotune presets
- How to download from baidu without account 2019
- How many people at john mayer where the light is
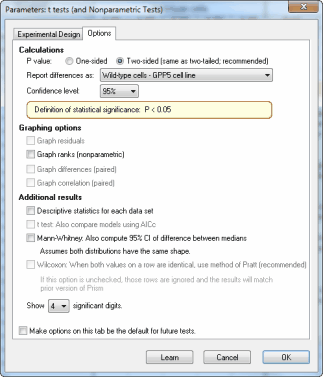
- Ks test graphpad prism 8
- Reason core security license key free
- Propane generator
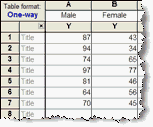
ZRF1 binds the H2A-ubiquitin mark catalyzed by the UV-RING1B complex, and its presence at damaged chromatin depends on the recognition factor XPC. We have recently shown that ZRF1 is an essential factor in NER. During NER H2A-ubiquitylation is catalyzed by the E3 ligase RNF8, the UV-DDB-CUL4 and UV-RING1B complexes. Furthermore, it was demonstrated that the RING1B-catalyzed ubiquitylation through Polycomb-repressive complex 1 (PRC1) mediates DSB-induced gene silencing, highlighting an additional function of H2A-ubiquitylation in DNA repair. At DSBs, ubiquitylation of H2A is carried out by the E3 ligases RNF168, RNF8, and RING1B, which facilitate signaling and accumulation of repair proteins. Although lesion-containing DNA associates with the nuclear matrix after UV irradiation it is less well understood how nuclear organization affects NER.Īnother important feature of DNA repair is H2A-ubiquitylation. Additionally, it has been recently shown that heterochromatin impedes CPD removal, and this process is enabled by DDB2. During NER, the damage recognition factor DDB2 promotes local chromatin decondensation and NER seems to involve large-scale chromatin rearrangements. In mammalian cells, nuclear organization during double-strand break (DSB) repair affects chromosome translocations and pathway choice. Studies of DNA repair have shown that nuclear positioning and migration of the damaged DNA to specific repair centers is a central component of many repair pathways. These complexes catalyze the mono-ubiquitylation of histones H2A, H3 and H4 as well as the polyubiquitylation of XPC.
DDB2 along with DDB1, the RING-domain proteins RBX1 or RING1B, and either of the scaffold proteins CUL4A or CUL4B forms E3 ubiquitin ligase complexes (UV-DDB-CUL4A/B and UV-RING1B). Efficient recognition of CPDs and 6-4 photoproducts also requires DDB2 (XPE). XPC specifically recognizes structures that distort the DNA double-helix, binds damaged DNA, and rapidly dissociates upon triggering NER. During GG-NER, lesions are detected by the damage recognition factors XPC and DDB2. In contrast, transcription-independent recognition of lesions is handled by global genome NER (GG-NER). Transcription-coupled NER (TC-NER) is limited to regions of active transcription, where RNA Polymerase II stalling elicits the DNA damage response. Mammalian NER recognizes DNA lesions by two different pathways.
#Ks test graphpad prism 8 skin
Defects in NER cause genetic disorders such as Xeroderma pigmentosum, which constitutes hypersensitivity to sunlight and a predisposition for skin cancer. Nucleotide excision repair (NER) is one of the major DNA repair pathways and handles various lesions such as cyclobutane pyrimidine dimers (CPDs) and 6-4 photoproducts, which occur after exposure to ultraviolet (UV) light. Our data thus provide insight into the sub-nuclear organization of NER and reveal a novel role for histone H2A-ubiquitylation. We further show that the H2A-ubiquitin binding protein ZRF1 resides in the nucleolus, and that it anchors ubiquitylated chromatin along with XPC. Employing a LacR-based tethering system we demonstrate that H2A-ubiquitylation via the UV-RING1B complex localizes chromatin close to the nucleolus. Upon inducing localized damage, we observe migration of damaged DNA towards the nucleolus. Analyzing unscheduled DNA synthesis (UDS) indicates that NER preferentially occurs in specific nuclear areas, viz the nucleolus.
Although lesion-containing DNA associates with the nuclear matrix after UV irradiation it is still not understood how nuclear organization affects NER. It is the primary pathway for repair of various DNA lesions caused by exposure to ultraviolet (UV) light, such as cyclobutane pyrimidine dimers (CPDs) and 6-4 photoproducts.

One of the major cellular DNA repair pathways is nucleotide excision repair (NER). Received: NovemAccepted: FebruPublished: MaAbstract Keywords: nucleotide excision repair, DNA repair, ubiquitin, chromatin, nuclear dynamics 1 Laboratory of Molecular Epigenetics, Institute of Molecular Biology, Mainz, Germany, Ackermannweg, Mainz, GermanyĢ Faculty of Biology, Johannes Gutenberg University, Mainz, Germany
